Welcome to HLRMDB
A comprehensive database of the human microbiome with metagenomic assembly, taxonomic classification and functional annotation by analysis of long-read and hybrid sequencing data.
The human microbiome harbours an immense diversity of uncultivated microbes; short-read metagenomic sequencing has elucidated much of this diversity, but fragment repeats and mobile elements constrain strain-level resolution. Fortunately, long-read metagenomic sequencing can generate reads spanning tens of kilobases with single-molecule accuracies more than 99%, enabling near-complete genome and gene cluster recovery in a cultivation-independent manner. However, systematic resources that aggregate and standardise such long-read outputs remain limited.
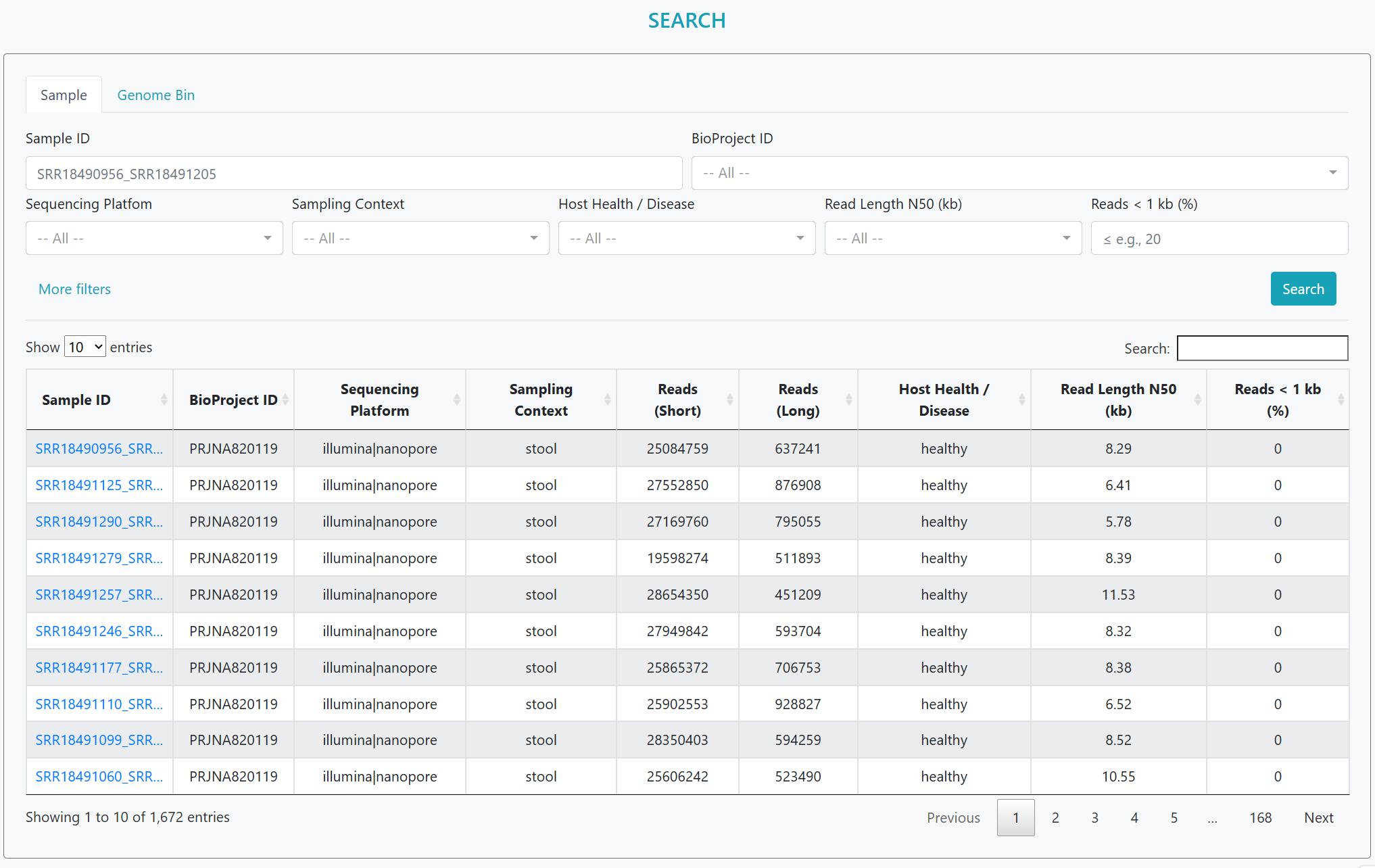
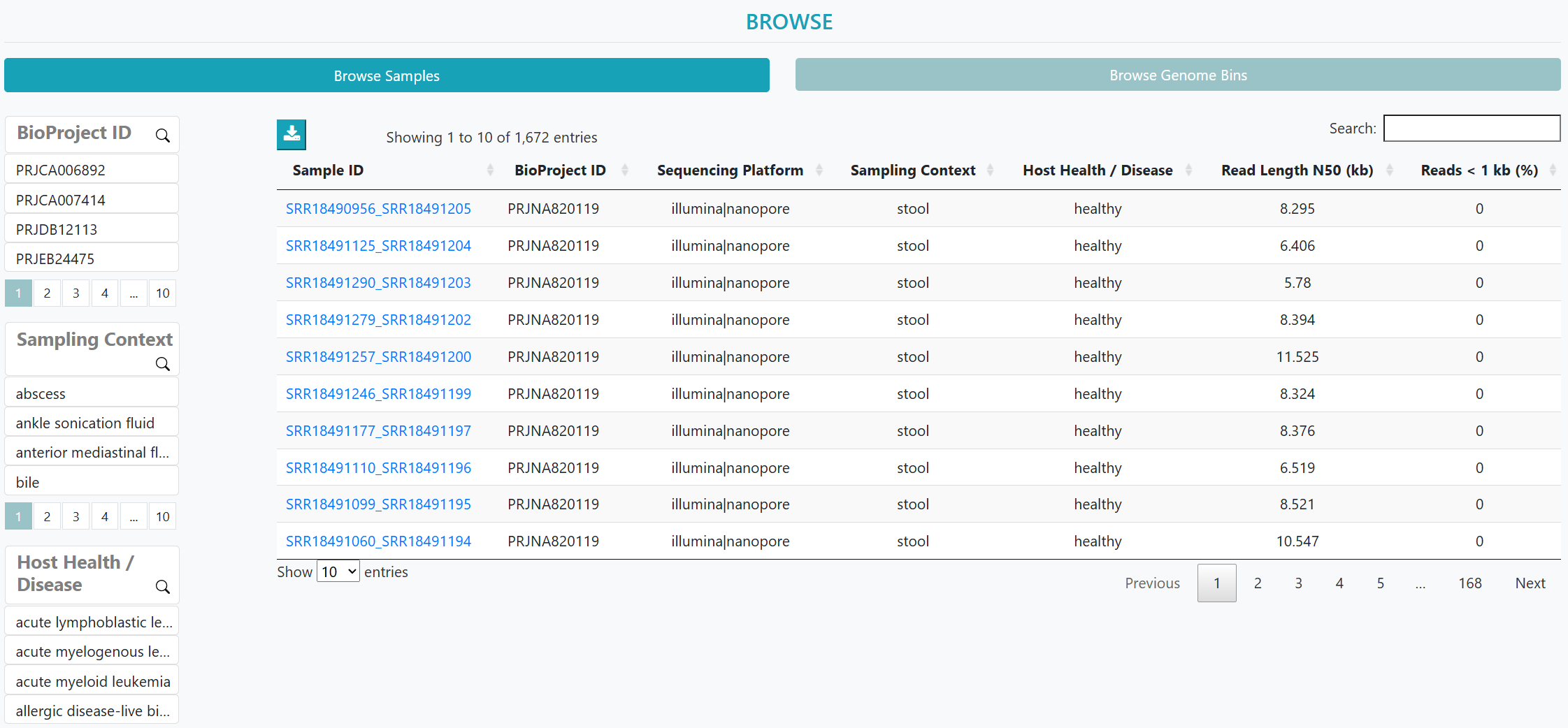
In this study, we curated 1,672 publicly available metagenomes (1,291 long reads; 381 hybrids) spanning 38 studies, 39 sampling contexts and 42 host health states.
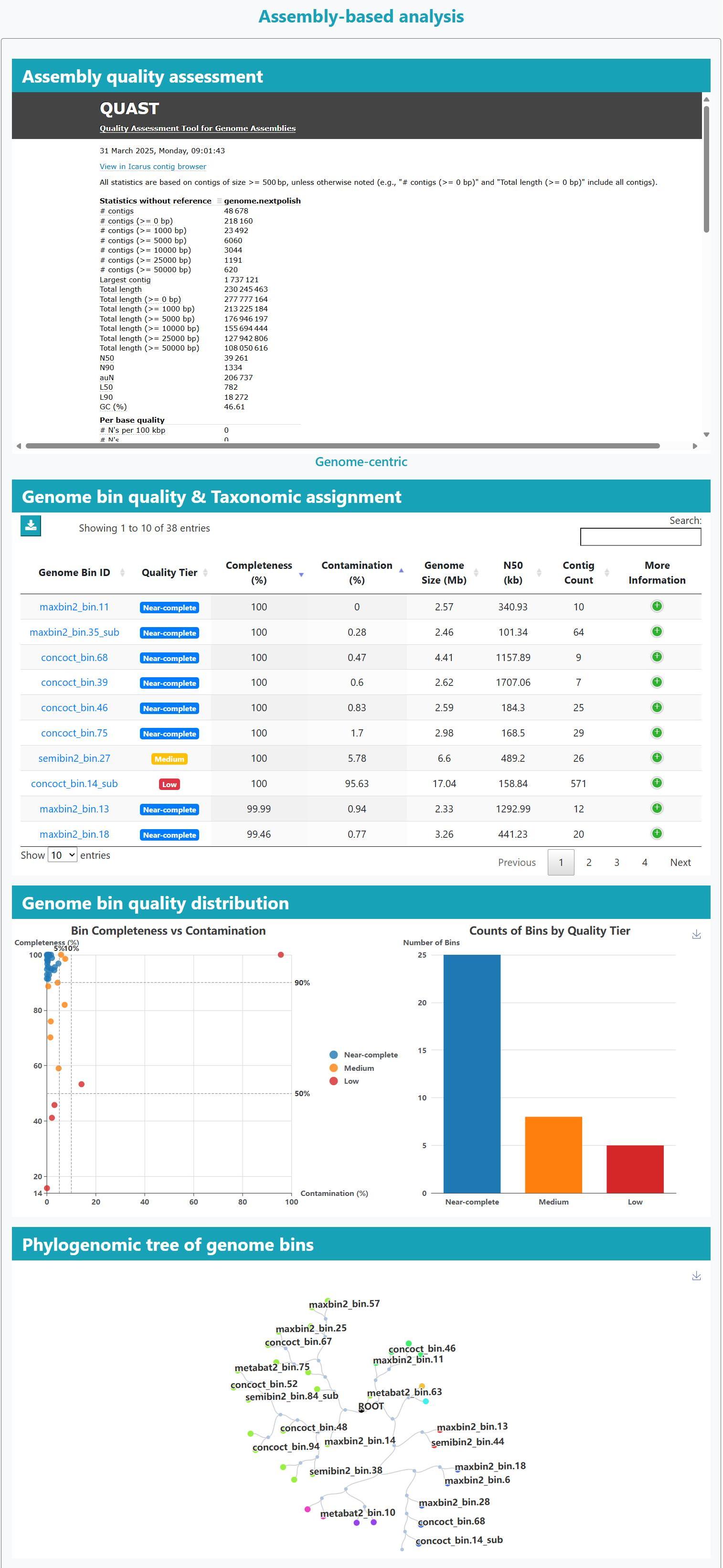
A uniform assembly and binning pipeline reconstructed > 98 Gb of contigs and yielded 18,721 genome bins, collectively spanning 21 phyla and 1,323 bacterial
species, with 6,339 meet near-complete quality standards, and an additional 5,609 are of medium quality. The remaining 6,773 bins are classified as low-quality drafts. HLRMDB integrates these genome-resolved data with extensive
gene-centric functional profiles and antimicrobial resistance annotations. An interactive web interface supports flexible access to both sample-level and
genome-level results, with multiple visualisations linking raw reads to assembled genomes. Overall, HLRMDB offers a harmonised, long-read-oriented
repository that supports reproducible, strain-resolved comparative genomics and context-sensitive ecological investigations of the human microbiome.
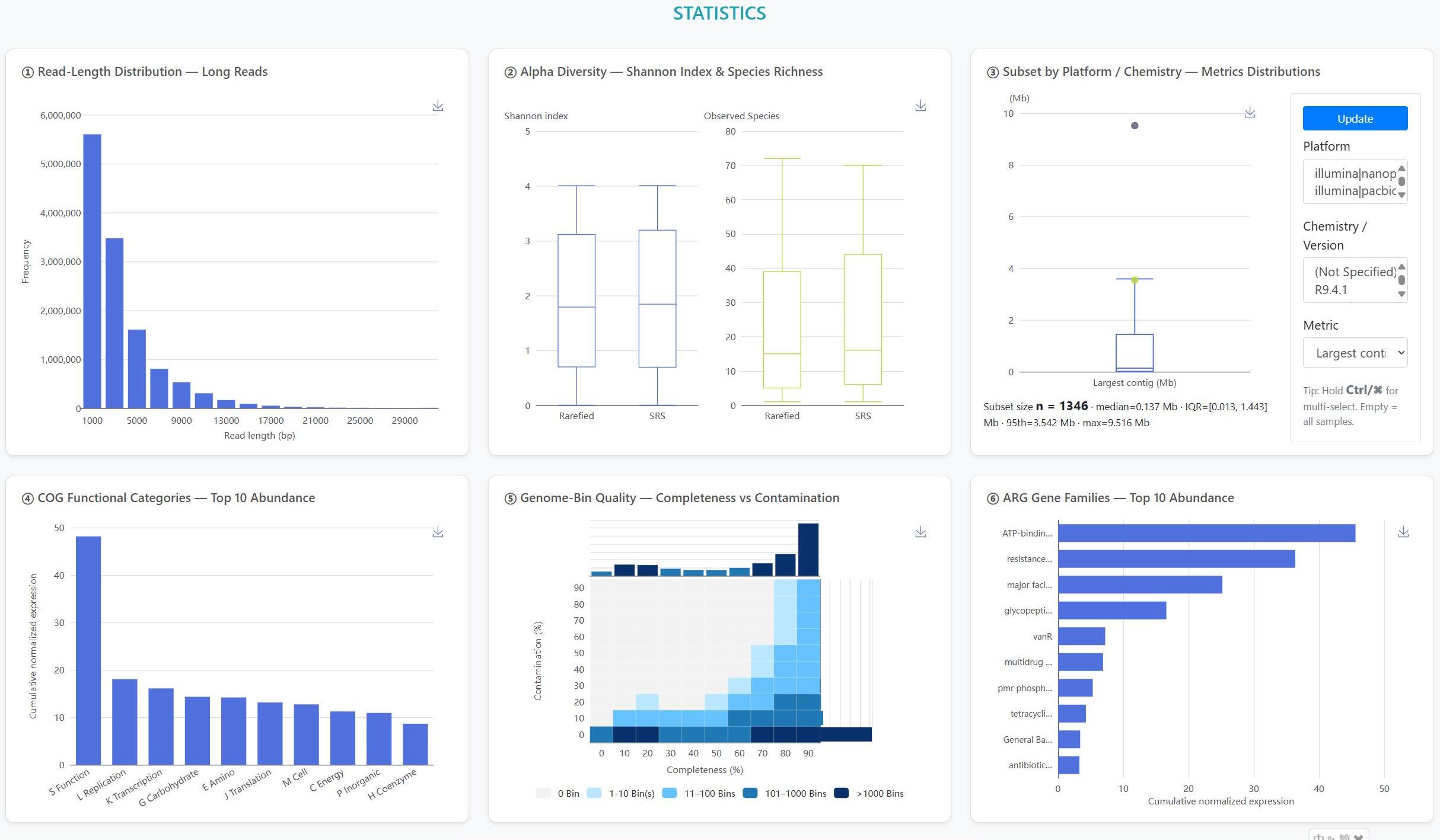
> Long-read focus: We curated 1,672 human-derived microbiome datasets, comprising 1,291 long-read datasets sequenced on third-generation platforms (1,290 ONT; 1 PacBio) and 381 hybrid datasets pairing Illumina short reads with long reads (357 ONT-hybrid; 24 PacBio-hybrid). The Statistics page now exposes Platform and Chemistry/Version filters (ONT: R7.3, R9.4, R9.4.1, R10.4.1/Kit14; PacBio: Sequel v3.0, RS II P6-C4), and flags small-n strata.
> Human provenance: All datasets derive exclusively from human-associated microbiome niches (e.g., stool, oral cavity, airway).
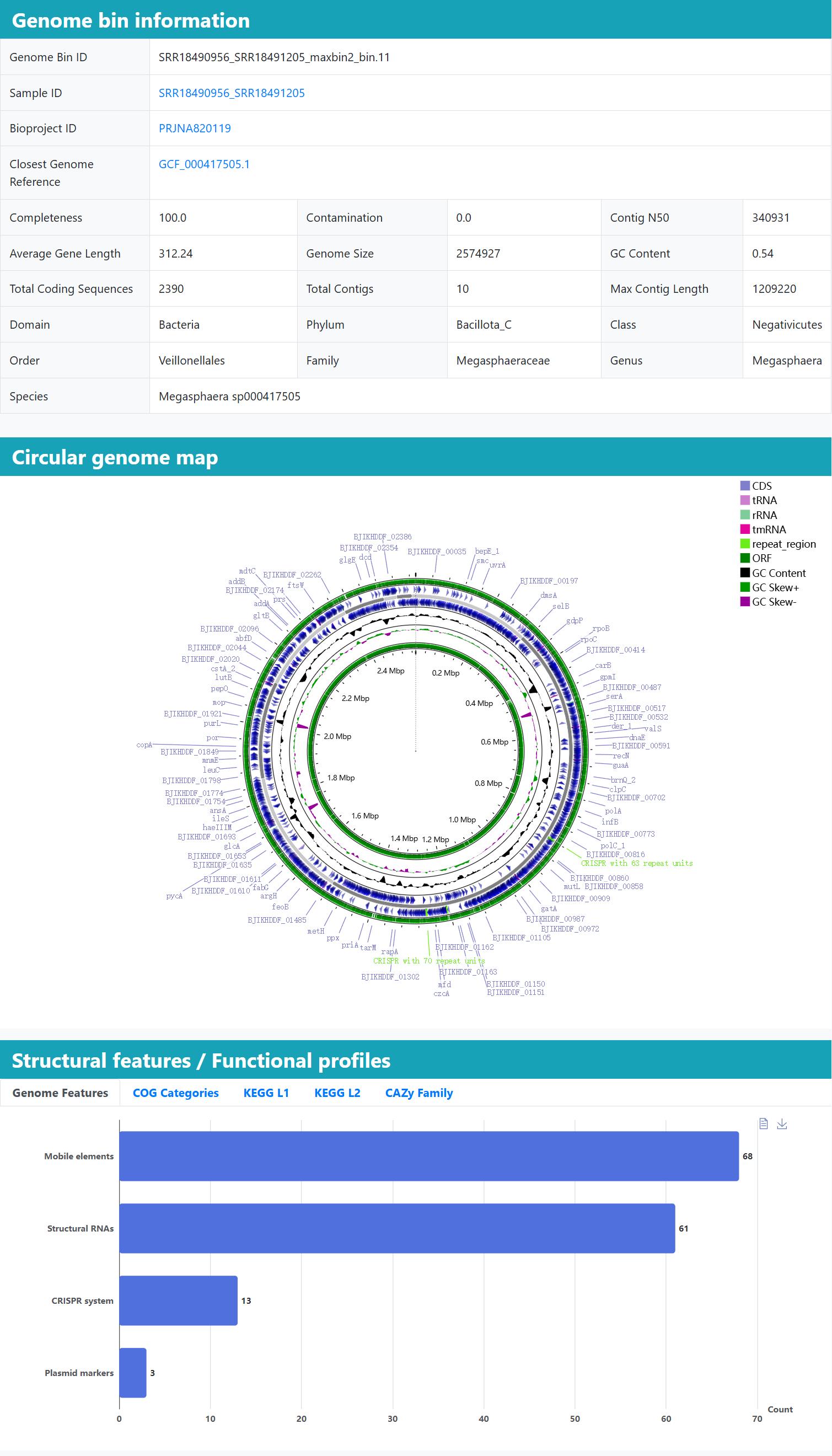
> Genome integrity: Long-read and hybrid assemblies deliver highly contiguous reconstructions; MAGs are often recovered as single- or few-contig genomes with higher completeness and markedly reduced fragmentation relative to short-read assemblies.
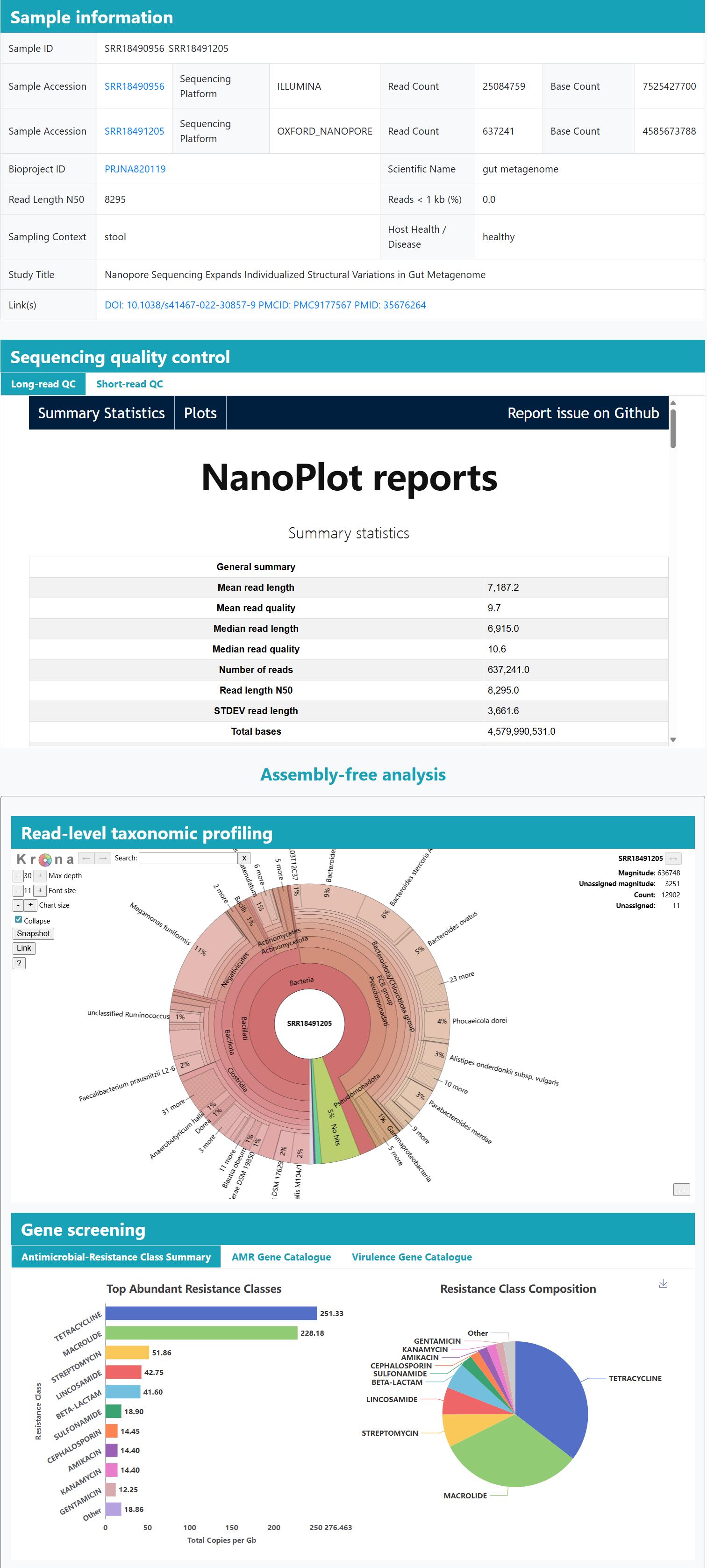
> Two-tier annotation: A two-tier framework couples genome-resolved gene and pathway catalogs from MAGs with read-based taxonomic and functional profiling, providing complementary assembly-based and assembly-free perspectives.
> Targeted retrieval: Sample- and Genome-centric search, faceted metadata filters, and dashboard-style exploration for rapid dataset triage.
> Community profiling: Interactive charts summarise each sample’s taxonomic composition, highlighting dominant and low-abundance lineages.
> Assembly quality: Each genome bin is accompanied by N50, completeness and contamination metrics for rapid quality assessment.
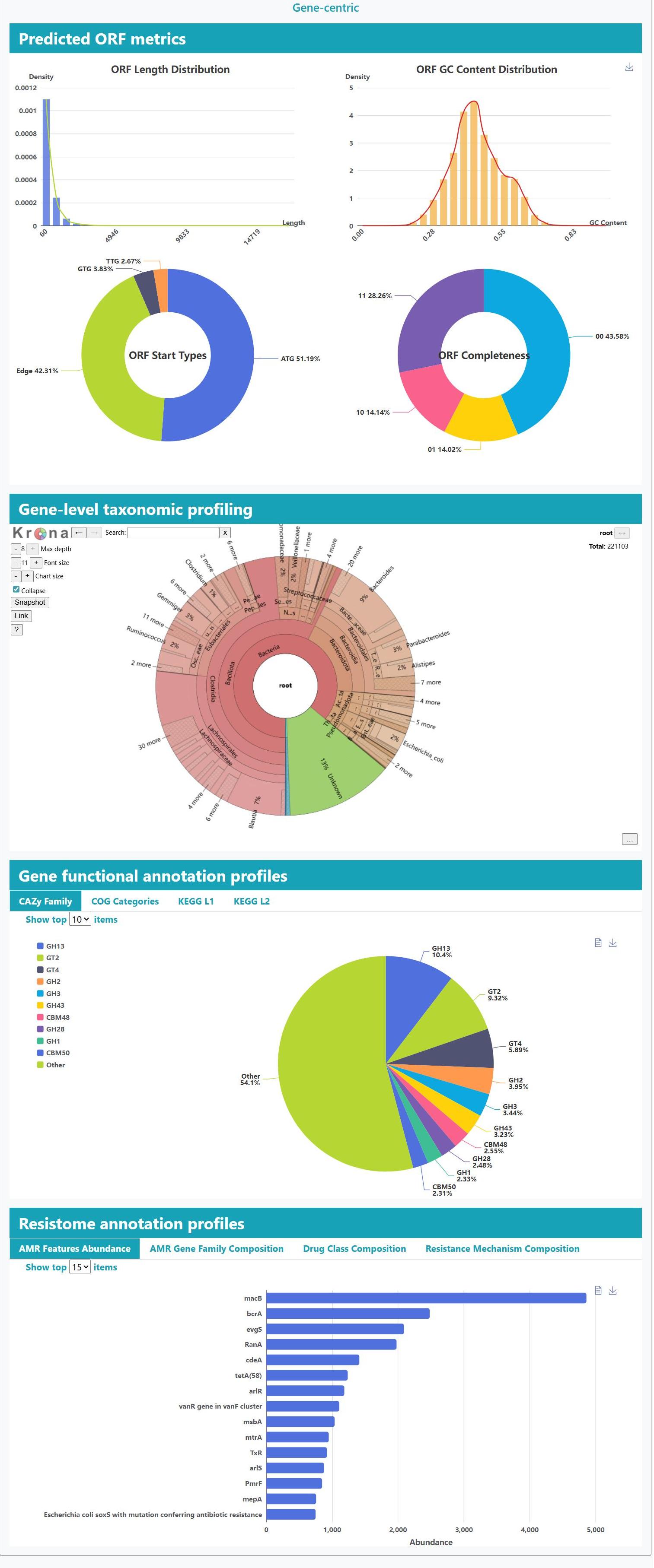
> Resistance context: Read- and assembly-level ARG profiling links hits to their contigs and MAG hosts, with interactive summaries by gene family, drug class, and mechanism.
> Multiscale function: Dynamic pie and bar plots for COG, KEGG and CAZy annotations, with interactive Top-N controls and on-the-fly database switching.
- HLRMDB v1.1 was released.
- HLRMDB v1.0 was released.
To get a better experience with this site, Chrome, Firefox are advised! RELATED RESOURCE: SRA | ENA | GSA
Contact us:HLRMDB is free only for academic usage. For commercial usage, please Contact Prof. Jianbo Pan, No. 1 Yixueyuan Road, Yuzhong District, Chongqing, 400016, P. R. China Email: panjianbo@cqmu.edu.cn