About
Genes profiles with temporal-specific expression patterns were defined as those whose expression has a specific pattern with respect to time.
Using a high-throughput regression analysis approach, eight linear and nonlinear parametric models (linear, constant, logistic, logarithmic, exponential, power, trigonometric, quadratic functions) were fitted to gene expression profiles from time-series experiments to identify eight types of genes with temporal-specific expression patterns. They serve as biological representations of diverse gene expression patterns.
We manually collected and curated raw data from 2684 time-series transcriptomic datasets in the Gene Expression Omnibus (GEO) and ArrayExpress repositories. These datasets were derived from well-conducted time course studies (that had at least five time points) involving humans and various other organisms.
For gene expression data, we clustered subsets of genes using Clust and performed regression analysis to identify patterns.
GeTeSEPdb Function Introduction
Fuzzy query is enabled. In the main page, time course datasets can be quickly searched by a single gene symbol or a keyword of dataset.
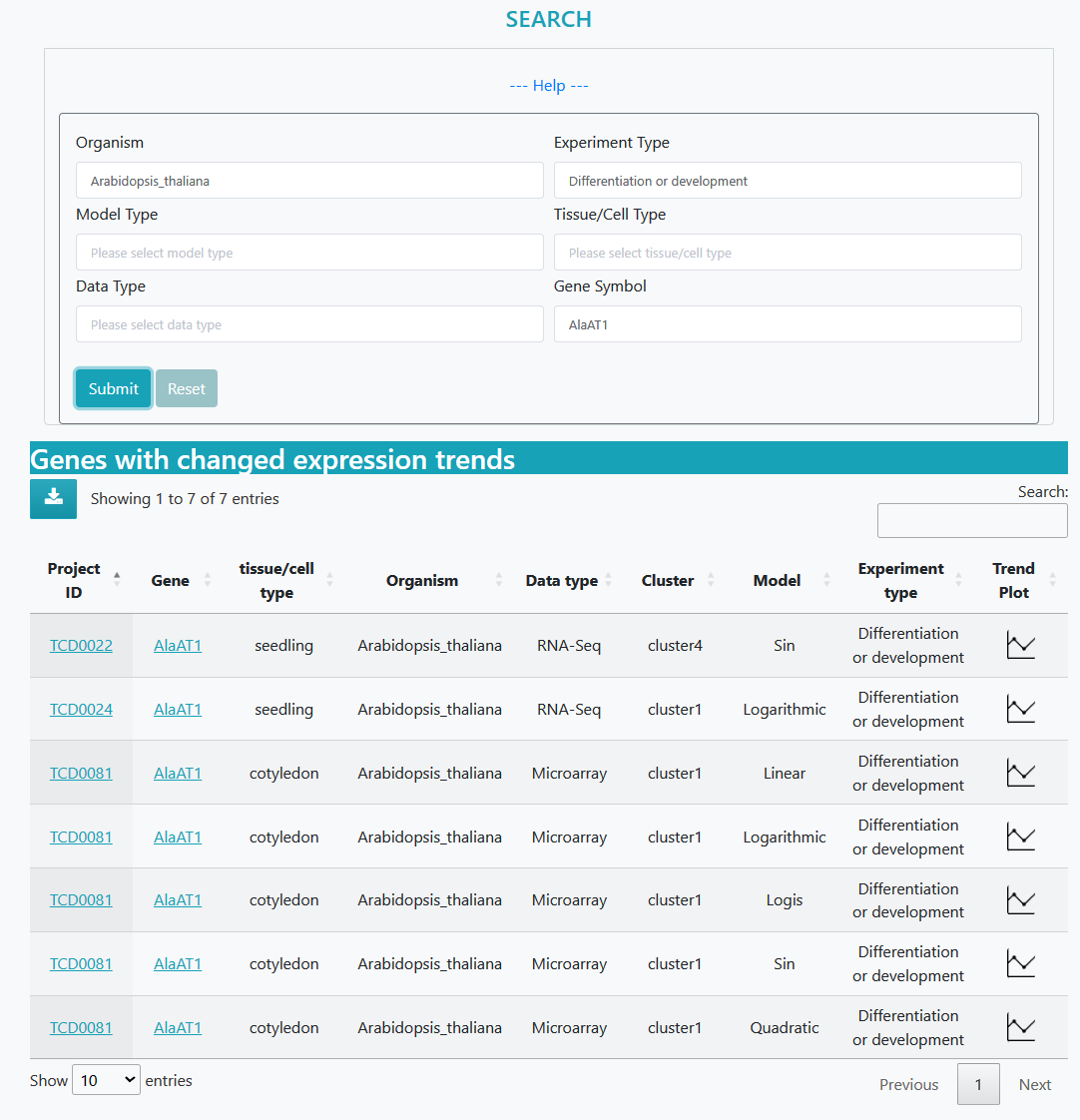
Search result page
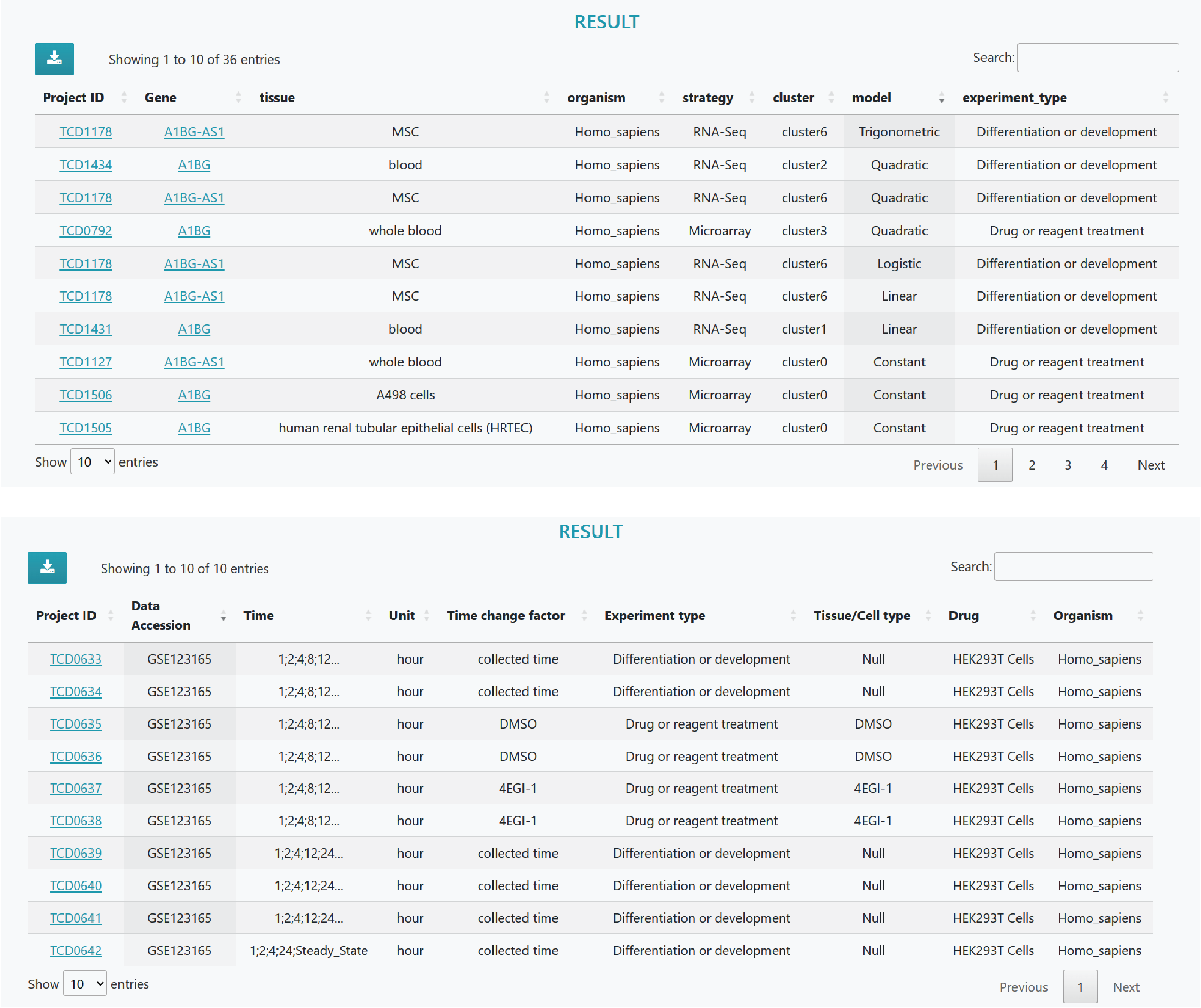
Time course datasets that meet the specified criteria are listed separately, and clicking on the dataset ID leads to a detailed information page about the study. Furthermore, clicking on a specific gene provides detailed information about that gene.
Study detailed information page
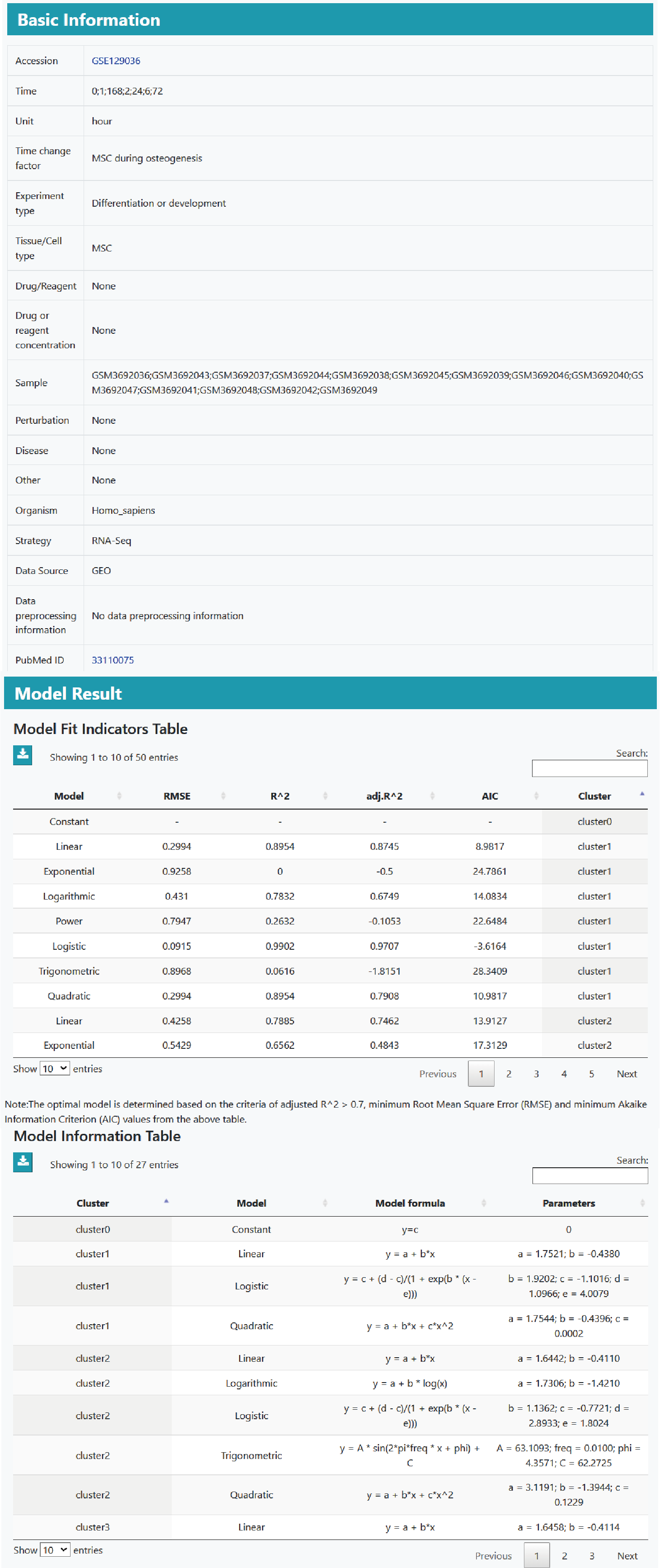
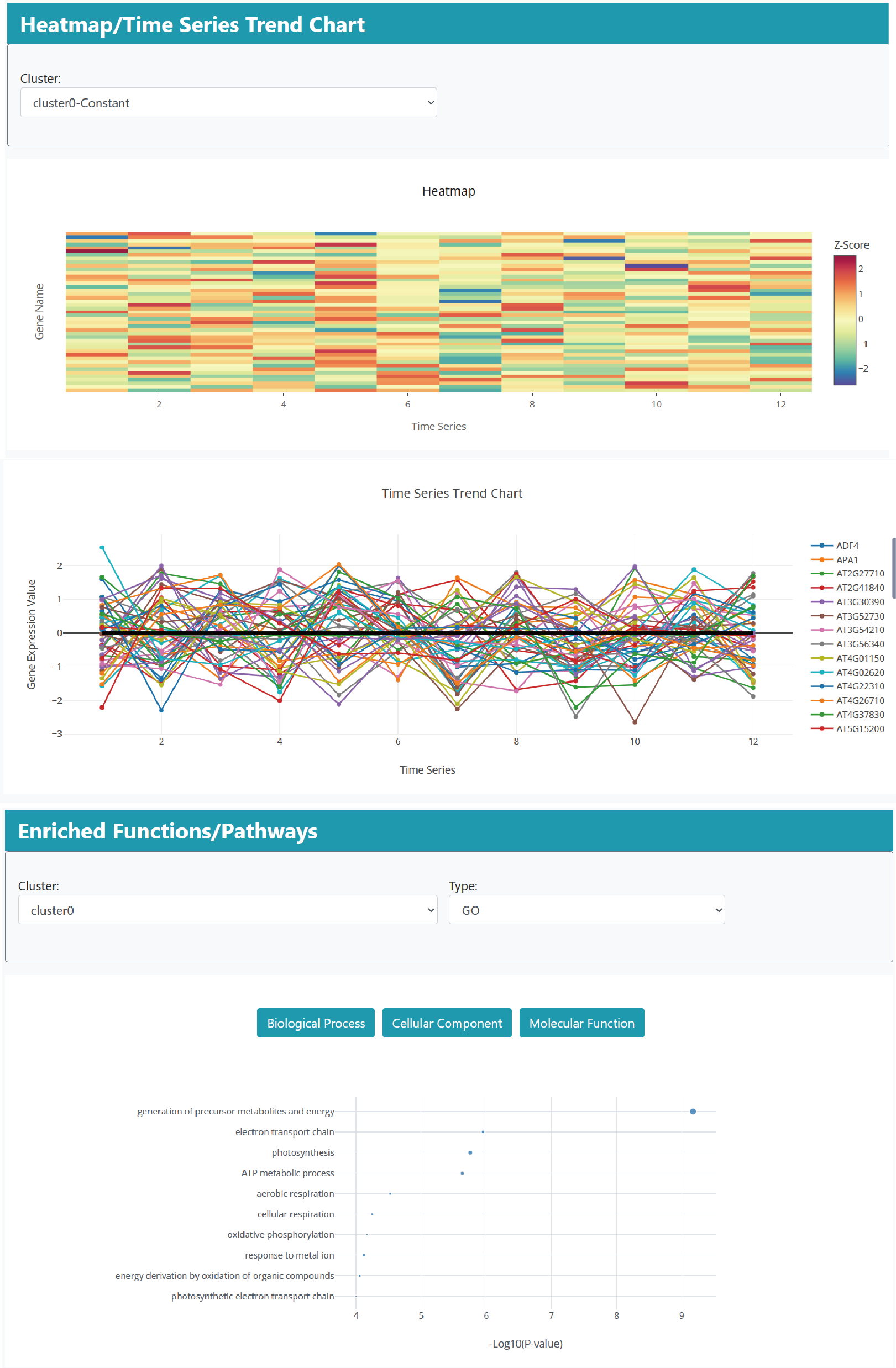
The detailed information page contains available experimental details, evaluation results of seven models, information about the optimal model, gene expression heat maps/trend maps, and functional annotation and pathway enrichment results for different model groups.
Gene detailed information page
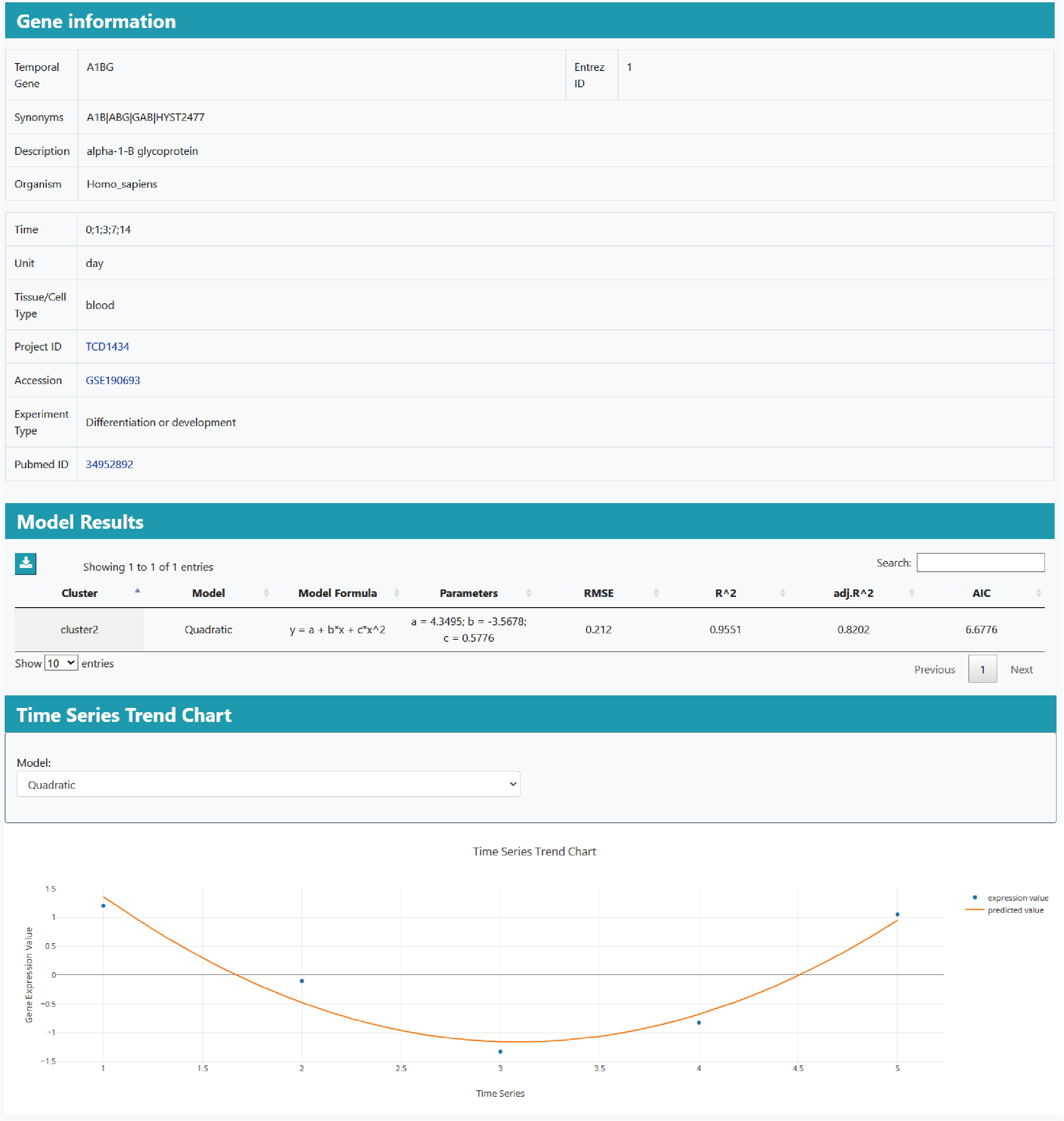
The detailed information page contains the available gene and experimental information, the details of the gene model, the heat map,trend map of gene expression and fitting.
By utilizing these parametric models, researchers can uncover diverse gene expression patterns, providing insights into regulatory mechanisms and functional implications in different biological contexts.
The first five conditions can be arbitrarily selected, and finally enter the gene you want to query, you can get the screening results.
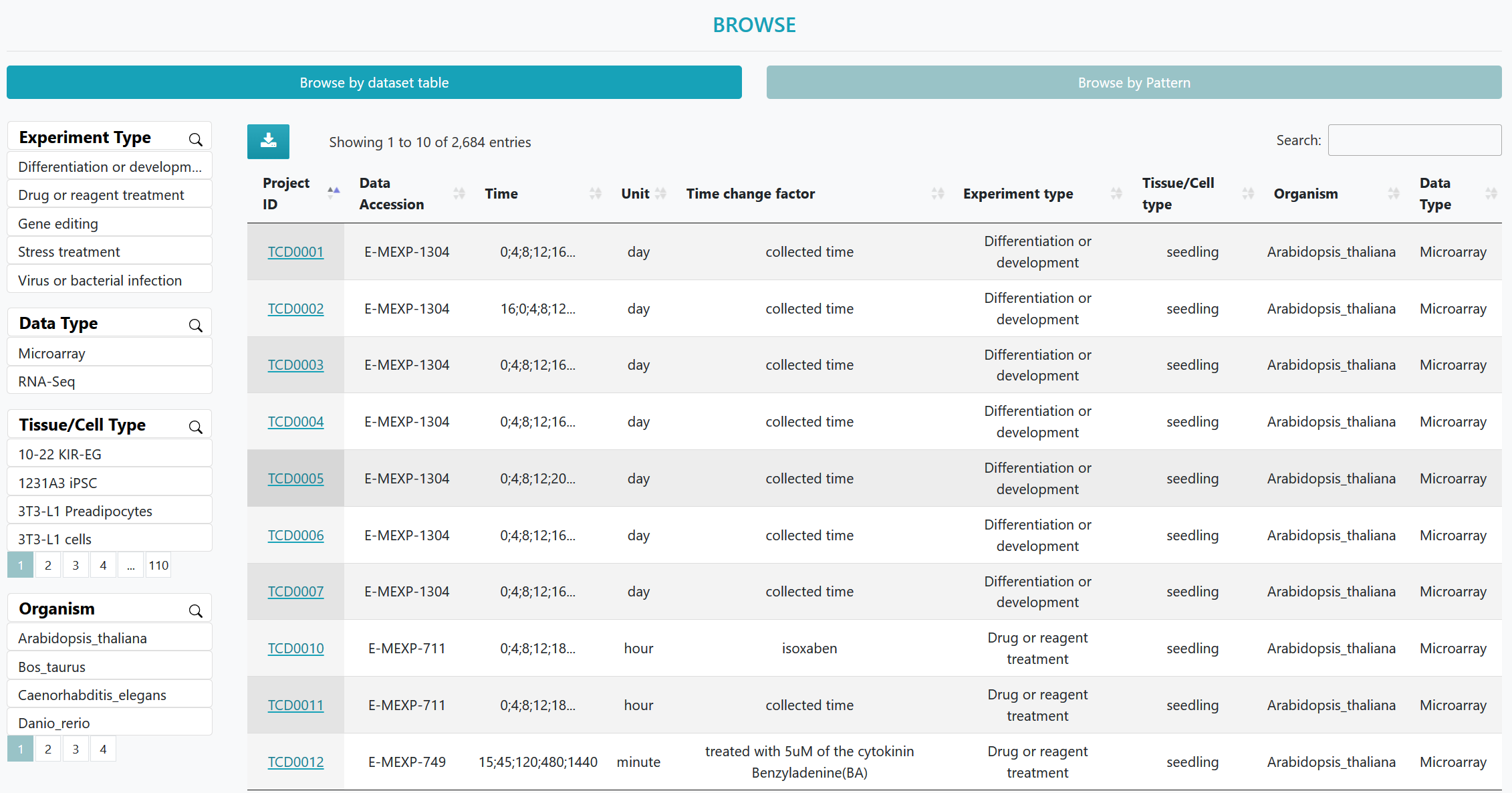
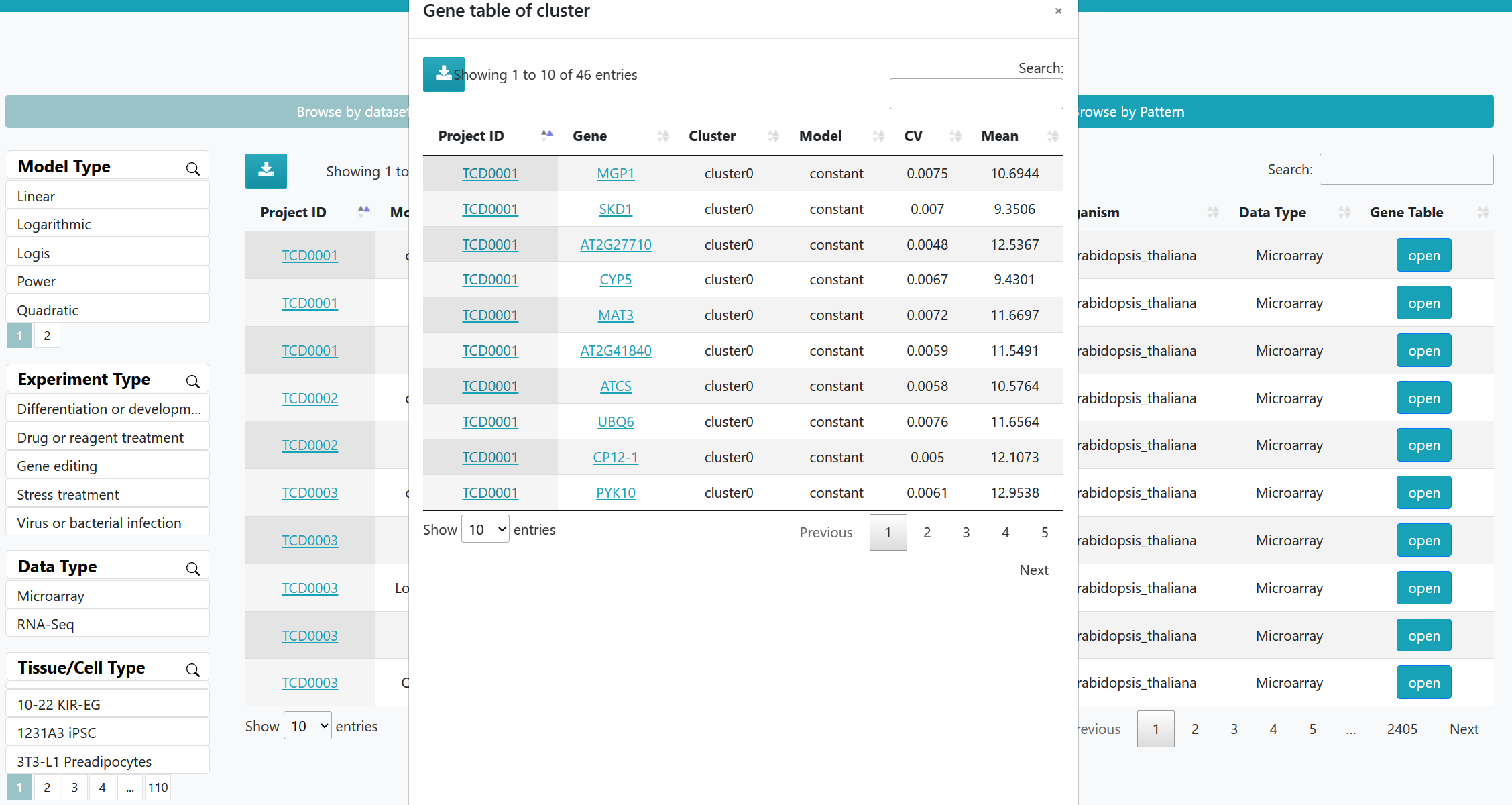
A list of time course datasets and pattern gene datasets can be viewed in an interactive table on the page. Users can customize filters based on "Data Type", "Tissue/Cell Type", and "Organism". Clicking on the "Dataset ID" and "Gene Name" allows access to the corresponding detailed information page.
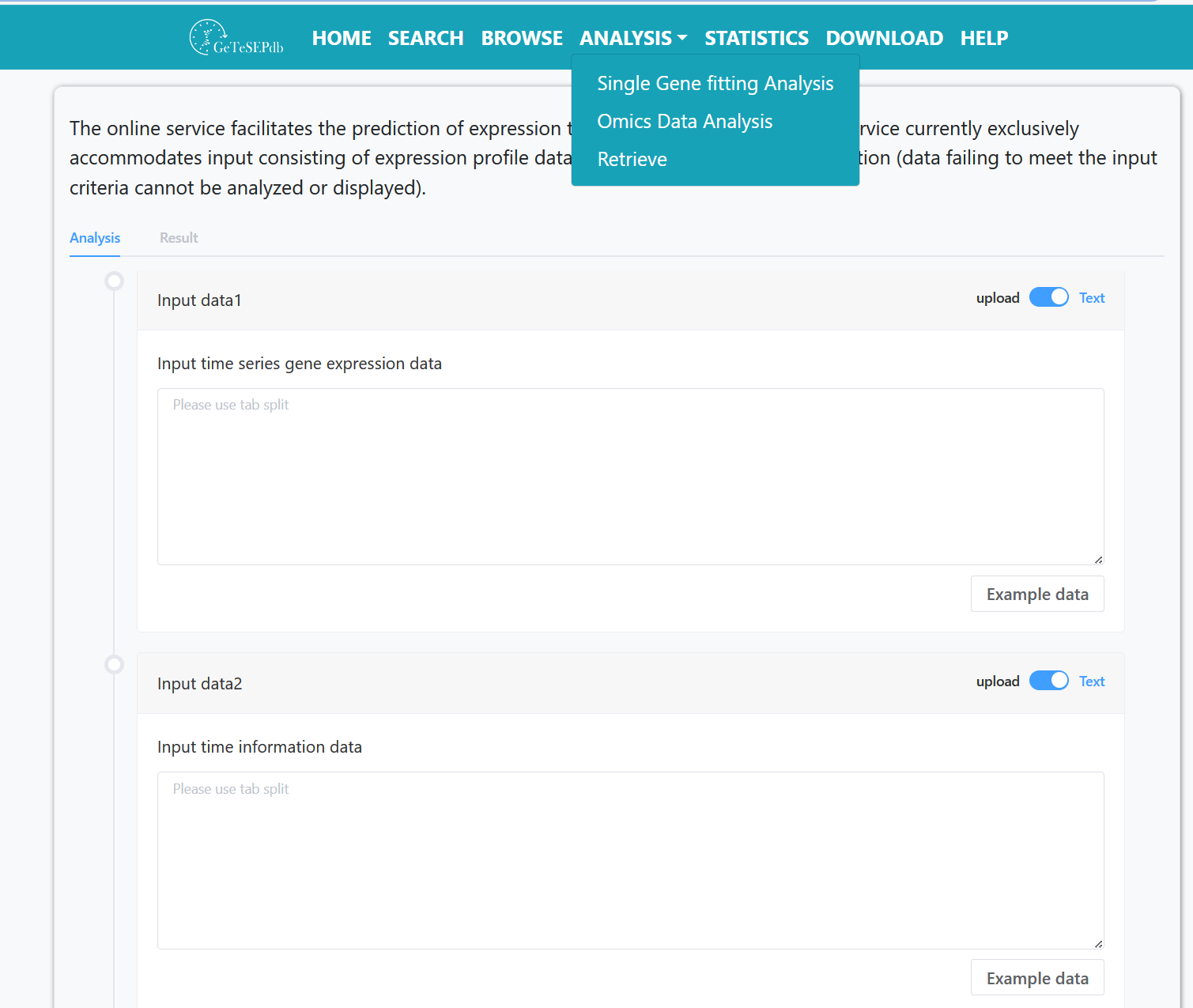
Classification results of gene expression patterns are obtained through Clust and regression analysis. When users upload correct gene expression data and corresponding sample time information files, analysis results are displayed and downloadable result files are provided.
Step 1: Upload time series gene expression data file.
Step 2: Upload sample information data file.
Step 3: Submit
Consequently, clustering and regression analysis yield classification results for gene expression patterns. Users can download compressed files containing all the results by clicking the "Download All Result" button.
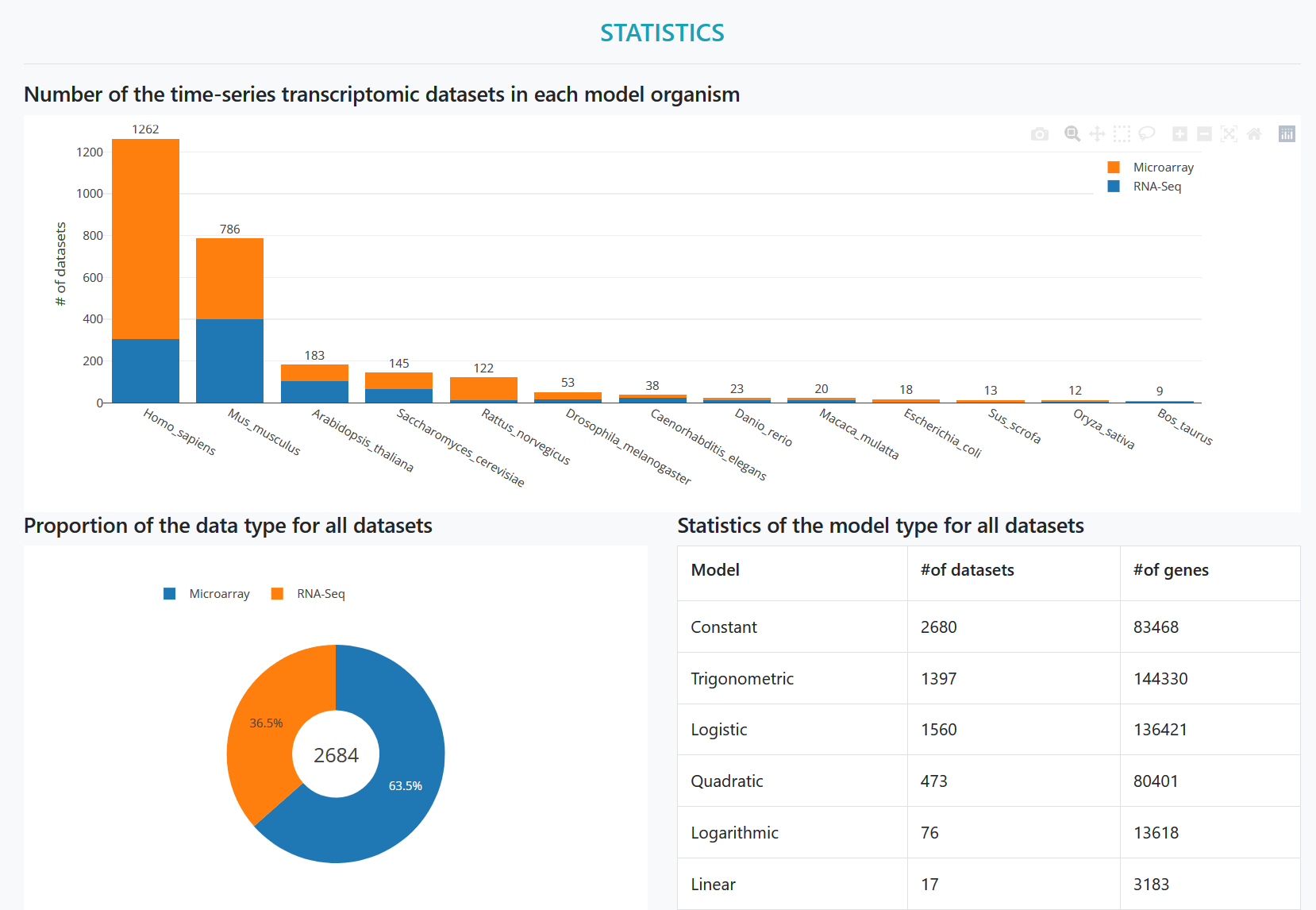
In the "Statistics" page, the statistics of the time course datasets in multiple perspectives are illustrated.
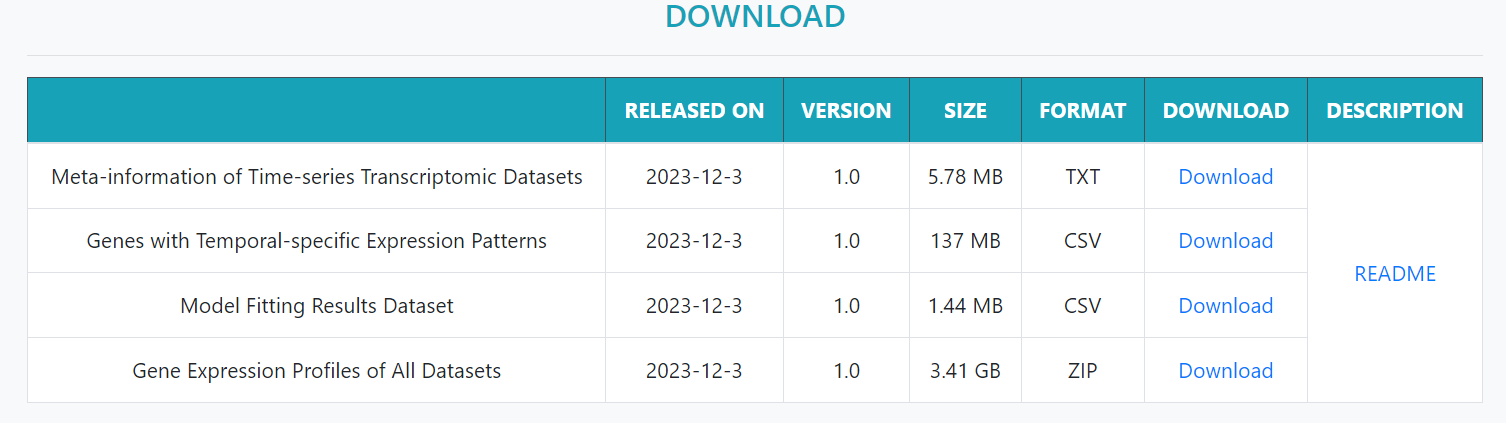
The Download page provides access to time course datasets information, available GeTeSEP data, model analysis results, and gene expression profiles for each dataset, all of which can be downloaded.
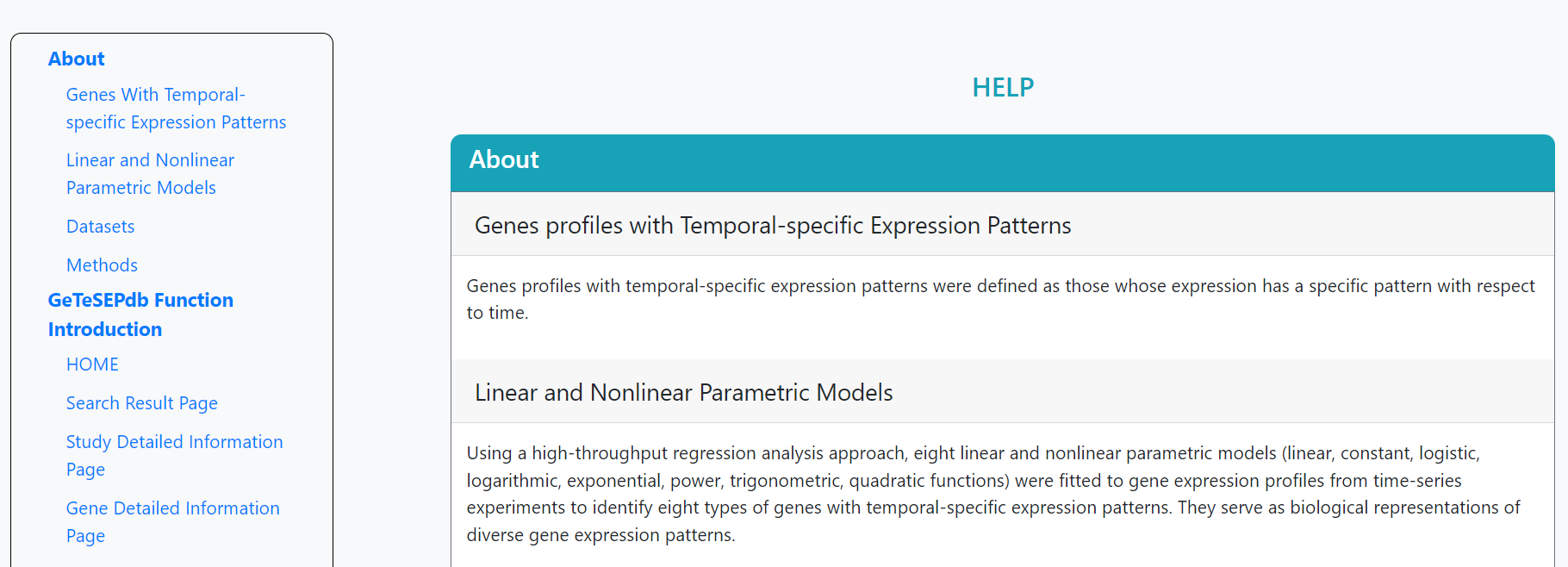

On the help page, the introduction of the features and the guidelines for the functions are stated. The contact details are attached at the bottom.
Contact Us
- Please feel free to contact Prof. Jianbo Pan with respect to any details pertaining to GeTeSEP.
-
>Address :
Center for Novel Target and Therapeutic Intervention, Institute of Life Sciences, Chongqing Medical University,
No. 1 Yixueyuan Road, Yuzhong District, Chongqing, 400016, P. R. China.
-
>Email :
panjianbo@cqmu.edu.cn